Smoothing With Backprop
If you’ve ever implemented forward-backward in an HMM (likely for a class assignment), you know this is an annoying exercise fraught with off-by-one errors or underflow issues.
A fun fact that has since been made concrete by Jason Eisner’s tutorial paper in 2016 is that backpropagation is forward-backward — if you implemented the forward pass for marginalisation for an HMM, then performing backpropagation will net you the result of forward-backward, or the smoothing result.
With auto-differentiation tools like pytorch gaining all the traction that it has, it’s a cheap and easy way to perform smoothing for structured prediction models like these (the result extends beyond HMMs and forward-backward).
Getting an intuition
Here, we lift an example from Sasha Rush’s PyTorch paper. Consider a simple Conditional Random Field (CRF):
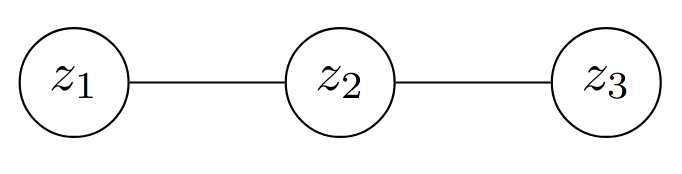
Each random variable $z_i$ can take on $C$ unique values, and so this model has $(N - 1) \times C \times C$ components (that’s how I’ll refer to these in this blogpost), where $N = 3$ and $N-1$ corresponds to the number of edges in the chain.
And so the energy of a specific configuration of this model when $z_1 = w, z_2 = x, z_3 = y$ ($w$, $x$, and $y$, are realisations of $z_i$ here) can be written as, $$\psi_{12}(w, x) \cdot \psi_{23}(x, y)$$ and $\psi$ here represents the edge potentials between each random variable.
The difficulty in these settings is usually in computing the probability, which requires you to normalise over all possible configurations or structures: \begin{align} \sum_{c_1,c_2,c_3 \in \mathcal{S}} \psi_{12}(c_1, c_2) \cdot \psi_{23}(c_2, c_3) \end{align} This set $\mathcal{S}$ contains all possible combinations of possible structures for which the naive computation would be to perform the full $C \times C \times C$ enumeration of $\mathcal{S}$. This is where the Forward Algorithm steps in to make life a little bit more polynomial, instead of exponential (and we really hate exponentials these days, amirite?).
So we know we can compute the energy per structure, and we know that we can find the sum of all structures (for these Linear Chain CRFs), in polynomial time.
More generally in structured probabilistic models (PCFG, HMM, Linear-chain CRF), we have this general form in the marginalisation:
\begin{align}
\log Z &= \log \sum_{j \in \mathrm{structures}} \exp\left( z_j \right) \\
z_j &= \sum_{k \in \mathrm{components}(j)} \psi_k
\end{align}
If we pick apart $\dfrac{\partial \log Z}{\partial \psi_k}$ with the chain rule:
\begin{align}
\dfrac{\partial \log Z}{\partial \psi_k} = \sum_{j’} \dfrac{\partial \log Z}{\partial z_{j’}} \dfrac{\partial z_{j’}}{\partial \psi_k}
\end{align}
we can first look at the derivative of log-sum-exp:
\begin{align}
\dfrac{\partial}{\partial z_{j’}} \log \sum_j \exp(z_j) = \frac{\exp(z_{j’})}{\sum_j \exp(z_j)}
\end{align}
This gives us softmax!
If you’re the sort more easily convinced by code:
z = torch.randn(10, requires_grad=True)
>>> L = torch.logsumexp(z, dim=-1)
>>> g, = torch.autograd.grad(L, z, retain_graph=True)
>>> g
tensor([0.0358, 0.0278, 0.1912, 0.0833, 0.0665, 0.1221, 0.1316, 0.1834, 0.0721,
0.0863])
>>> torch.softmax(z, dim=-1)
tensor([0.0358, 0.0278, 0.1912, 0.0833, 0.0665, 0.1221, 0.1316, 0.1834, 0.0721,
0.0863], grad_fn=<SoftmaxBackward>)
How do we interpret this softmax? This is the probability over the latent structure. $z_j$ is the log-potentials over the individual structure, and the derivative here are the potentials normalised over all of the possible structures.
Returning to our simple CRF example, consider how that maps to this general form. A specific $j$-th structure maps to an assignment of $c_1, c_2, c_3$. More concretely, we can say something like: \begin{align} z_{c_1, c_2, c_3} = \log \psi_{12}(c_1, c_2) + \log \psi_{23}(c_2, c_3) \end{align}
Now given a fixed assignment of $c_2$ and $c_3$, the assignment of $c_1$ can vary, but perhaps we would like to know:
What is the probability that the edge $c_2 = x$ and $c_3 = y$ exists, among all the possible structures?
Naively, we’d sum up the energy for all the structures where the assignments hold true, \begin{align} \frac{ \sum_{c_1, c_2, c_3\in \mathcal{S} | c_2 = x, c_3 = y} \psi_{12}(c_1, x) \cdot \psi_{23}(x, y)}{ \sum_{c_1,c_2,c_3 \in \mathcal{S}} \psi_{12}(c_1, c_2) \cdot \psi_{23}(c_2, c_3)} \end{align} However, if we knew the probabilities for each structure, we could sum up the probabilities for the ones where $c_2 = x$ and $c_3 = y$ hold.
Now let us consider $\dfrac{\partial z_{j’}}{\partial \psi_k}$, and in our current example, $\psi_k = \log \psi_{23}(x, y)$. Only the structures with that component present will be $1$, while every other structure will be $0$.
If we compute the chain rule as in Equation (4), we recover the marginal over that particular component!
Ok great. What’s this for.
In the case of CRFs or HMM, performing the Forward Algorithm, and then running backprop with respect to the log probabilities or log-potentials, will give you the Forward-backward algorithm, for free!
Like I mentioned before, CRFs and HMMs aren’t the only place where this trick applies. Sasha Rush has a framework called torch-struct that has a collection of different algorithms related to structured prediction. He goes further…
Replacing log-sum-exp with $\max$
Or as Sasha Rush calls it: semirings.
To recover Viterbi, swap out all instances of $\log \sum \exp$ before with $\max$. The resulting gradients will give you the $\mathrm{argmax}$ of where the optimal category should be, all without having to separately implement Viterbi with backtracking.
>>> L_max = torch.max(z, dim=-1)[0]
>>> g_max, = torch.autograd.grad(L_max, z, retain_graph=True)
>>> g_max
tensor([0., 0., 1., 0., 0., 0., 0., 0., 0., 0.])
>>> torch.argmax(z)
tensor(2)
Closing arguments
I think the significance of Eisner’s tutorial may not have been as well appreciated at the time due to the prevalence of libraries that perform these structured prediction tasks. Moreover, automatic differentiation libraries weren’t as popular. If a library already exists that does forward-backward, viterbi, inside-outside, etc. nicely for you, why bother implementing it again with some fancy backprop trick that you have to roll yourself?
These days, these algorithms are a little less trendy, but autodifferentiation libraries are everywhere. They’re not completely gone though. If you look closely enough, you’ll find structured prediction in Connectionist Temporal Classification, which is still being used for speech recognition.
If nothing else, it’s a shortcut for completing your NLP assignment.
@misc{tan2021-03-14,
title = {Smoothing With Backprop},
author = {Tan, Shawn},
howpublished = {\url{https://blog.wtf.sg/posts/2021-03-14-smoothing-with-backprop/}},
year = {2021}
}